Nengo frontend API¶
Nengo Objects¶
A network contains ensembles, nodes, connections, and other networks. |
|
A group of neurons that collectively represent a vector. |
|
An interface for making connections directly to an ensemble’s neurons. |
|
Provide non-neural inputs to Nengo objects and process outputs. |
|
Connects two objects together. |
|
An interface for making connections to a learning rule. |
|
A probe is an object that collects data from the simulation. |
-
class
nengo.Network(label=None, seed=None, add_to_container=None)[source]¶ A network contains ensembles, nodes, connections, and other networks.
A network is primarily used for grouping together related objects and connections for visualization purposes. However, you can also use networks as a nice way to reuse network creation code.
To group together related objects that you do not need to reuse, you can create a new
Networkand add objects in awithblock. For example:network = nengo.Network() with network: with nengo.Network(label="Vision"): v1 = nengo.Ensemble(n_neurons=100, dimensions=2) with nengo.Network(label="Motor"): sma = nengo.Ensemble(n_neurons=100, dimensions=2) nengo.Connection(v1, sma)
To reuse a group of related objects, you can create a new subclass of
Network, and add objects in the__init__method. For example:class OcularDominance(nengo.Network): def __init__(self): self.column = nengo.Ensemble(n_neurons=100, dimensions=2) network = nengo.Network() with network: left_eye = OcularDominance() right_eye = OcularDominance() nengo.Connection(left_eye.column, right_eye.column)
- Parameters
- labelstr, optional
Name of the network.
- seedint, optional
Random number seed that will be fed to the random number generator. Setting the seed makes the network’s build process deterministic.
- add_to_containerbool, optional
Determines if this network will be added to the current container. If None, this network will be added to the network at the top of the
Network.contextstack unless the stack is empty.
- Attributes
- connectionslist
Connectioninstances in this network.- ensembleslist
Ensembleinstances in this network.- labelstr
Name of this network.
- networkslist
Networkinstances in this network.- nodeslist
Nodeinstances in this network.- probeslist
Probeinstances in this network.- seedint
Random seed used by this network.
-
property
all_objects¶ (list) All objects in this network and its subnetworks.
-
property
all_ensembles¶ (list) All ensembles in this network and its subnetworks.
-
property
all_nodes¶ (list) All nodes in this network and its subnetworks.
-
property
all_networks¶ (list) All networks in this network and its subnetworks.
-
property
all_connections¶ (list) All connections in this network and its subnetworks.
-
property
all_probes¶ (list) All probes in this network and its subnetworks.
-
property
n_neurons¶ (int) Number of neurons in this network, including subnetworks.
-
class
nengo.Ensemble(*args, **kwargs)[source]¶ A group of neurons that collectively represent a vector.
- Parameters
- n_neuronsint
The number of neurons.
- dimensionsint
The number of representational dimensions.
- radiusint, optional
The representational radius of the ensemble.
- encodersDistribution or (n_neurons, dimensions) array_like, optional
The encoders used to transform from representational space to neuron space. Each row is a neuron’s encoder; each column is a representational dimension.
- interceptsDistribution or (n_neurons,) array_like, optional
The point along each neuron’s encoder where its activity is zero. If
eis the neuron’s encoder, then the activity will be zero whendot(x, e) <= c, wherecis the given intercept.- max_ratesDistribution or (n_neurons,) array_like, optional
The activity of each neuron when the input signal
xis magnitude 1 and aligned with that neuron’s encodere; i.e., whendot(x, e) = 1.- eval_pointsDistribution or (n_eval_points, dims) array_like, optional
The evaluation points used for decoder solving, spanning the interval (-radius, radius) in each dimension, or a distribution from which to choose evaluation points.
- n_eval_pointsint, optional
The number of evaluation points to be drawn from the
eval_pointsdistribution. If None, then a heuristic is used to determine the number of evaluation points.- neuron_type
NeuronType, optional The model that simulates all neurons in the ensemble (see
NeuronType).- gainDistribution or (n_neurons,) array_like
The gains associated with each neuron in the ensemble. If None, then the gain will be solved for using
max_ratesandintercepts.- biasDistribution or (n_neurons,) array_like
The biases associated with each neuron in the ensemble. If None, then the gain will be solved for using
max_ratesandintercepts.- noiseProcess, optional
Random noise injected directly into each neuron in the ensemble as current. A sample is drawn for each individual neuron on every simulation step.
- normalize_encodersbool, optional
Indicates whether the encoders should be normalized.
- labelstr, optional
A name for the ensemble. Used for debugging and visualization.
- seedint, optional
The seed used for random number generation.
- Attributes
- biasDistribution or (n_neurons,) array_like or None
The biases associated with each neuron in the ensemble.
- dimensionsint
The number of representational dimensions.
- encodersDistribution or (n_neurons, dimensions) array_like
The encoders, used to transform from representational space to neuron space. Each row is a neuron’s encoder, each column is a representational dimension.
- eval_pointsDistribution or (n_eval_points, dims) array_like
The evaluation points used for decoder solving, spanning the interval (-radius, radius) in each dimension, or a distribution from which to choose evaluation points.
- gainDistribution or (n_neurons,) array_like or None
The gains associated with each neuron in the ensemble.
- interceptsDistribution or (n_neurons) array_like or None
The point along each neuron’s encoder where its activity is zero. If
eis the neuron’s encoder, then the activity will be zero whendot(x, e) <= c, wherecis the given intercept.- labelstr or None
A name for the ensemble. Used for debugging and visualization.
- max_ratesDistribution or (n_neurons,) array_like or None
The activity of each neuron when
dot(x, e) = 1, whereeis the neuron’s encoder.- n_eval_pointsint or None
The number of evaluation points to be drawn from the
eval_pointsdistribution. If None, then a heuristic is used to determine the number of evaluation points.- n_neuronsint or None
The number of neurons.
- neuron_typeNeuronType
The model that simulates all neurons in the ensemble (see
nengo.neurons).- noiseProcess or None
Random noise injected directly into each neuron in the ensemble as current. A sample is drawn for each individual neuron on every simulation step.
- radiusint
The representational radius of the ensemble.
- seedint or None
The seed used for random number generation.
-
property
neurons¶ A direct interface to the neurons in the ensemble.
-
property
size_in¶ The dimensionality of the ensemble.
-
property
size_out¶ The dimensionality of the ensemble.
-
class
nengo.ensemble.Neurons(ensemble)[source]¶ An interface for making connections directly to an ensemble’s neurons.
This should only ever be accessed through the
neuronsattribute of an ensemble, as a way to signal toConnectionthat the connection should be made directly to the neurons rather than to the ensemble’s decoded value, e.g.:with nengo.Network(): a = nengo.Ensemble(10, 1) b = nengo.Ensemble(10, 1) nengo.Connection(a.neurons, b.neurons)
-
property
ensemble¶ (Ensemble) The ensemble these neurons are part of.
-
property
probeable¶ (tuple) Signals that can be probed in the neuron population.
-
property
size_in¶ (int) The number of neurons in the population.
-
property
size_out¶ (int) The number of neurons in the population.
-
property
-
class
nengo.Node(*args, **kwargs)[source]¶ Provide non-neural inputs to Nengo objects and process outputs.
Nodes can accept input, and perform arbitrary computations for the purpose of controlling a Nengo simulation. Nodes are typically not part of a brain model per se, but serve to summarize the assumptions being made about sensory data or other environment variables that cannot be generated by a brain model alone.
Nodes can also be used to test models by providing specific input signals to parts of the model, and can simplify the input/output interface of a
Networkwhen used as a relay to/from its internal ensembles (seeEnsembleArrayfor an example).- Parameters
- outputcallable, array_like, or None
Function that transforms the Node inputs into outputs, a constant output value, or None to transmit signals unchanged.
- size_inint, optional
The number of dimensions of the input data parameter.
- size_outint, optional
The size of the output signal. If None, it will be determined based on the values of
outputandsize_in.- labelstr, optional
A name for the node. Used for debugging and visualization.
- seedint, optional
The seed used for random number generation. Note: no aspects of the node are random, so currently setting this seed has no effect.
- Attributes
- labelstr
The name of the node.
- outputcallable, array_like, or None
The given output.
- size_inint
The number of dimensions for incoming connection.
- size_outint
The number of output dimensions.
-
class
nengo.Connection(*args, **kwargs)[source]¶ Connects two objects together.
The connection between the two object is unidirectional, transmitting information from the first argument,
pre, to the second argument,post.Almost any Nengo object can act as the pre or post side of a connection. Additionally, you can use Python slice syntax to access only some of the dimensions of the pre or post object.
For example, if
nodehassize_out=2andensemblehassize_in=1:with nengo.Network() as net: node = nengo.Node(np.zeros(2)) ensemble = nengo.Ensemble(10, 1)
We could not create the following connection:
with net: nengo.Connection(node, ensemble)
But, we could create either of these two connections:
with net: nengo.Connection(node[0], ensemble) nengo.Connection(node[1], ensemble)
- Parameters
- preEnsemble or Neurons or Node
The source Nengo object for the connection.
- postEnsemble or Neurons or Node or LearningRule
The destination object for the connection.
- synapseSynapse or None, optional
Synapse model to use for filtering (see
Synapse). If None, no synapse will be used and information will be transmitted without any delay (if supported by the backend—some backends may introduce a single time step delay).Note that at least one connection must have a synapse that is not None if components are connected in a cycle. Furthermore, a synaptic filter with a zero time constant is different from a None synapse as a synaptic filter will always add a delay of at least one time step.
- functioncallable or (n_eval_points, size_mid) array_like, optional
Function to compute across the connection. Note that
premust be an ensemble to apply a function across the connection. If an array is passed, the function is implicitly defined by the points in the array and the providedeval_points, which have a one-to-one correspondence.- transform(size_out, size_mid) array_like, optional
Linear transform mapping the pre output to the post input. This transform is in terms of the sliced size; if either pre or post is a slice, the transform must be shaped according to the sliced dimensionality. Additionally, the function is applied before the transform, so if a function is computed across the connection, the transform must be of shape
(size_out, size_mid).- solverSolver, optional
Solver instance to compute decoders or weights (see
Solver). Ifsolver.weightsis True, a full connection weight matrix is computed instead of decoders.- learning_rule_typeLearningRuleType or iterable of LearningRuleType, optional
Modifies the decoders or connection weights during simulation.
- eval_points(n_eval_points, size_in) array_like or int, optional
Points at which to evaluate
functionwhen computing decoders, spanning the interval (-pre.radius, pre.radius) in each dimension. If None, will use the eval_points associated withpre.- scale_eval_pointsbool, optional
Indicates whether the evaluation points should be scaled by the radius of the pre Ensemble.
- labelstr, optional
A descriptive label for the connection.
- seedint, optional
The seed used for random number generation.
- Attributes
- functioncallable
The given function.
- function_sizeint
The output dimensionality of the given function. If no function is specified, function_size will be 0.
- labelstr
A human-readable connection label for debugging and visualization. If not overridden, incorporates the labels of the pre and post objects.
- learning_rule_typeinstance or list or dict of LearningRuleType, optional
The learning rule types.
- postEnsemble or Neurons or Node or Probe or ObjView
The given post object.
- post_objEnsemble or Neurons or Node or Probe
The underlying post object, even if
postis anObjView.- post_sliceslice or list or None
The slice associated with
postif it is an ObjView, or None.- preEnsemble or Neurons or Node or ObjView
The given pre object.
- pre_objEnsemble or Neurons or Node
The underlying pre object, even if
postis anObjView.- pre_sliceslice or list or None
The slice associated with
preif it is an ObjView, or None.- seedint
The seed used for random number generation.
- solverSolver
The Solver instance that will be used to compute decoders or weights (see
nengo.solvers).- synapseSynapse
The Synapse model used for filtering across the connection (see
nengo.synapses).- transform(size_out, size_mid) array_like
Linear transform mapping the pre function output to the post input.
-
property
learning_rule¶ (LearningRule or iterable) Connectable learning rule object(s).
-
property
size_in¶ (int) The number of output dimensions of the pre object.
Also the input size of the function, if one is specified.
-
property
size_mid¶ (int) The number of output dimensions of the function, if specified.
If the function is not specified, then
size_in == size_mid.
-
property
size_out¶ (int) The number of input dimensions of the post object.
Also the number of output dimensions of the transform.
-
class
nengo.connection.LearningRule(connection, learning_rule_type)[source]¶ An interface for making connections to a learning rule.
Connections to a learning rule are to allow elements of the network to affect the learning rule. For example, learning rules that use error information can obtain that information through a connection.
Learning rule objects should only ever be accessed through the
learning_ruleattribute of a connection.-
property
connection¶ (Connection) The connection modified by the learning rule.
-
property
modifies¶ (str) The variable modified by the learning rule.
-
property
probeable¶ (tuple) Signals that can be probed in the learning rule.
-
property
size_out¶ (int) Cannot connect from learning rules, so always 0.
-
property
-
class
nengo.Probe(*args, **kwargs)[source]¶ A probe is an object that collects data from the simulation.
This is to be used in any situation where you wish to gather simulation data (spike data, represented values, neuron voltages, etc.) for analysis.
Probes do not directly affect the simulation.
All Nengo objects can be probed (except Probes themselves). Each object has different attributes that can be probed. To see what is probeable for each object, print its
probeableattribute.with nengo.Network(): ens = nengo.Ensemble(10, 1) print(ens.probeable)
('decoded_output', 'input', 'scaled_encoders')- Parameters
- targetEnsemble, Neurons, Node, or Connection
The object to probe.
- attrstr, optional
The signal to probe. Refer to the target’s
probeablelist for details. If None, the first element in theprobeablelist will be used.- sample_everyfloat, optional
Sampling period in seconds. If None, the
dtof the simluation will be used.- synapseSynapse, optional
A synaptic model to filter the probed signal.
- solverSolver, optional
Solverto compute decoders for probes that require them.- labelstr, optional
A name for the probe. Used for debugging and visualization.
- seedint, optional
The seed used for random number generation.
- Attributes
- attrstr or None
The signal that will be probed. If None, the first element of the target’s
probeablelist will be used.- sample_everyfloat or None
Sampling period in seconds. If None, the
dtof the simluation will be used.- solverSolver or None
Solverto compute decoders. Only used for probes of an ensemble’s decoded output.- synapseSynapse or None
A synaptic model to filter the probed signal.
- targetEnsemble, Neurons, Node, or Connection
The object to probe.
-
property
obj¶ (Nengo object) The underlying Nengo object target.
-
property
size_in¶ (int) Dimensionality of the probed signal.
-
property
size_out¶ (int) Cannot connect from probes, so always 0.
-
property
slice¶ (slice) The slice associated with the Nengo object target.
Distributions¶
A base class for probability distributions. |
|
Convenience function to sample a distribution or return samples. |
|
An arbitrary distribution from a PDF. |
|
A uniform distribution. |
|
A Gaussian distribution. |
|
An exponential distribution (optionally with high values clipped). |
|
Uniform distribution on or in an n-dimensional unit hypersphere. |
|
Sequence for quasi Monte Carlo sampling the |
|
Quasirandom distribution over the hypersphere or hyperball. |
|
Discrete distribution across a set of possible values. |
|
A set of samples. |
|
Distribution of the square root of a Beta distributed random variable. |
|
Distribution of the length of a subvectors of a unit vector. |
|
Distribution of the cosine of the angle between two random vectors. |
-
class
nengo.dists.Distribution[source]¶ A base class for probability distributions.
The only thing that a probabilities distribution need to define is a
Distribution.samplemethod. This base class ensures that all distributions accept the same arguments for the sample function.-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
-
class
nengo.dists.DistributionParam(name, default=Unconfigurable, optional=False, readonly=None)[source]¶ A Distribution.
-
class
nengo.dists.DistOrArrayParam(name, default=Unconfigurable, sample_shape=None, sample_dtype=<class 'numpy.float64'>, optional=False, readonly=None)[source]¶ Can be a Distribution or samples from a distribution.
-
nengo.dists.get_samples(dist_or_samples, n, d=None, rng=numpy.random)[source]¶ Convenience function to sample a distribution or return samples.
Use this function in situations where you accept an argument that could be a distribution, or could be an
array_likeof samples.- Parameters
- dist_or_samples
Distributionor (n, d) array_like Source of the samples to be returned.
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return.
- rngRandomState, optional
Random number generator.
- dist_or_samples
- Returns
- samples(n, d) array_like
Examples
from nengo.dists import get_samples rng = np.random.RandomState(seed=0) def mean(values, n=100): samples = get_samples(values, n=n, rng=rng) print(f"{np.mean(samples):.4f}") mean([1, 2, 3, 4]) mean(nengo.dists.Gaussian(0, 1))
2.5000 0.0598
-
class
nengo.dists.PDF(x, p)[source]¶ An arbitrary distribution from a PDF.
- Parameters
- xvector_like (n,)
Values of the points to sample from (interpolated).
- pvector_like (n,)
Probabilities of the
xpoints.
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.Uniform(low, high, integer=False)[source]¶ A uniform distribution.
It’s equally likely to get any scalar between
lowandhigh.Note that the order of
lowandhighdoesn’t matter; iflow < highthis will still work, andlowwill still be a closed interval whilehighis open.- Parameters
- lowNumber
The closed lower bound of the uniform distribution; samples >= low
- highNumber
The open upper bound of the uniform distribution; samples < high
- integerboolean, optional
If true, sample from a uniform distribution of integers. In this case, low and high should be integers.
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.Gaussian(mean, std)[source]¶ A Gaussian distribution.
This represents a bell-curve centred at
meanand with spread represented by the standard deviation,std.- Parameters
- meanNumber
The mean of the Gaussian.
- stdNumber
The standard deviation of the Gaussian.
- Raises
- ValidationError if std is <= 0
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.Exponential(scale, shift=0.0, high=inf)[source]¶ An exponential distribution (optionally with high values clipped).
If
highis left to its default value of infinity, this is a standard exponential distribution. Ifhighis set, then any sampled values at or abovehighwill be clipped so they are slightly belowhigh. This is useful for thresholding.The probability distribution function (PDF) is given by:
| 0 if x < shift p(x) = | 1/scale * exp(-(x - shift)/scale) if x >= shift and x < high | n if x == high - eps | 0 if x >= high
where
nis such that the PDF integrates to one, andepsis an infinitesimally small number such that samples ofxare strictly less thanhigh(in practice,epsdepends on floating point precision).- Parameters
- scalefloat
The scale parameter (inverse of the rate parameter lambda). Larger values make the distribution narrower (sharper peak).
- shiftfloat, optional
Amount to shift the distribution by. There will be no values smaller than this shift when sampling from the distribution.
- highfloat, optional
All values larger than or equal to this value will be clipped to slightly less than this value.
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.UniformHypersphere(surface=False, min_magnitude=0)[source]¶ Uniform distribution on or in an n-dimensional unit hypersphere.
Sample points are uniformly distributed across the volume (default) or surface of an n-dimensional unit hypersphere.
- Parameters
- surfacebool, optional
Whether sample points should be distributed uniformly over the surface of the hyperphere (True), or within the hypersphere (False).
- min_magnitudeNumber, optional
Lower bound on the returned vector magnitudes (such that they are in the range
[min_magnitude, 1]). Must be in the range [0, 1). Ignored ifsurfaceisTrue.
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.QuasirandomSequence[source]¶ Sequence for quasi Monte Carlo sampling the
[0, 1]-cube.This is similar to
np.random.uniform(0, 1, size=(num, d)), but with the additional property that eachd-dimensional point is uniformly scattered.While the sequence is defined deterministically, we introduce two stochastic elements to encourage heterogeneity in models using these sequences. First, we offset the start of the sequence by a random number between 0 and 1 to ensure we don’t oversample points aligned to the step size. Second, we shuffle the resulting sequence before returning to ensure we don’t introduce correlations between parameters sampled from this distribution.
This is based on the tutorial and code from 1.
See also
References
- 1
Martin Roberts. “The Unreasonable Effectiveness of Quasirandom Sequences.” https://web.archive.org/web/20211029094035/http://extremelearning.com.au/unreasonable-effectiveness-of-quasirandom-sequences/
Examples
rd = nengo.dists.QuasirandomSequence().sample(10000, 2) plt.scatter(*rd.T, c=np.arange(len(rd)), cmap='Blues', s=7)
-
sample(self, n, d=1, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.ScatteredHypersphere(surface=False, min_magnitude=0, base=QuasirandomSequence(), method='sct-approx')[source]¶ Quasirandom distribution over the hypersphere or hyperball.
Applies a spherical transform to the given quasirandom sequence (by default
QuasirandomSequence) to obtain uniformly scattered samples.This distribution has the nice mathematical property that the discrepancy between the empirical distribution and \(n\) samples is \(\widetilde{\mathcal{O}} (1 / n)\) as opposed to \(\mathcal{O} (1 / \sqrt{n})\) for the Monte Carlo method [1]. This means that the number of samples is effectively squared, making this useful as a means for sampling
eval_pointsandencoders.- Parameters
- surfacebool, optional
Whether sample points should be distributed uniformly over the surface of the hyperphere (True), or within the hypersphere (False).
- min_magnitudeNumber, optional
Lower bound on the returned vector magnitudes (such that they are in the range
[min_magnitude, 1]). Must be in the range [0, 1). Ignored ifsurfaceisTrue.- base
Distribution, optional The base distribution from which to sample quasirandom numbers.
- method{“sct-approx”, “sct”, “tfww”}
Method to use for mapping points to the hypersphere.
“sct-approx”: Same as “sct”, but uses lookup table to approximate the beta distribution, making it faster with almost exactly the same result.
“sct”: Use the exact Spherical Coordinate Transform (section 1.5.2 of [1]).
“tfww”: Use the Tashiro-Fang-Wang-Wong method (section 4.3 of [1]). Faster than “sct” and “sct-approx”, with the same level of uniformity for larger numbers of samples (
n >= 4000, approximately).
See also
Notes
The
QuasirandomSequencedistribution is mostly deterministic. Nondeterminism comes from a randomd-dimensional rotation.References
Examples
Plot points sampled from the surface of the sphere in 3 dimensions:
from mpl_toolkits.mplot3d import Axes3D points = nengo.dists.ScatteredHypersphere(surface=True).sample(1000, d=3) ax = plt.subplot(111, projection="3d") ax.scatter(*points.T, s=5)
Plot points sampled from the volume of the sphere in 2 dimensions (i.e. circle):
points = nengo.dists.ScatteredHypersphere(surface=False).sample(1000, d=2) plt.scatter(*points.T, s=5)
-
classmethod
spherical_transform_sct(samples, approx=False)[source]¶ Map samples from the
[0, 1]-cube onto the hypersphere.Uses the SCT method described in section 1.5.3 of Fang and Wang (1994).
-
static
spherical_transform_tfww(c_samples)[source]¶ Map samples from the
[0, 1]-cube onto the hypersphere surface.Uses the TFWW method described in section 4.3 of Fang and Wang (1994).
-
sample(self, n, d=1, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.Choice(options, weights=None)[source]¶ Discrete distribution across a set of possible values.
The same as Numpy random’s
choice, except can take vector or matrix values for the choices.- Parameters
- options(N, …) array_like
The options (choices) to choose between. The choice is always done along the first axis, so if
optionsis a matrix, the options are the rows of that matrix.- weights(N,) array_like, optional
Weights controlling the probability of selecting each option. Will automatically be normalized. If None, weights be uniformly distributed.
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.Samples(samples)[source]¶ A set of samples.
This class is a subclass of
Distributionso that it can be used in any situation that calls for aDistribution. However, the call toDistribution.samplemust match the dimensions of the samples or aValidationErrorwill be raised.- Parameters
- samples(n, d) array_like
nanddmust match what is eventually passed to
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
class
nengo.dists.SqrtBeta(n, m=1)[source]¶ Distribution of the square root of a Beta distributed random variable.
Given
n + mdimensional random unit vectors, the length of subvectors withmelements will be distributed according to this distribution.- Parameters
- n: int
Number of subvectors.
- m: int, optional
Length of each subvector.
See also
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
cdf(x)[source]¶ Cumulative distribution function.
Note
Requires SciPy.
- Parameters
- xarray_like
Evaluation points in [0, 1].
- Returns
- cdfarray_like
Probability that
X <= x.
-
class
nengo.dists.SubvectorLength(dimensions, subdimensions=1)[source]¶ Distribution of the length of a subvectors of a unit vector.
- Parameters
- dimensionsint
Dimensionality of the complete unit vector.
- subdimensionsint, optional
Dimensionality of the subvector.
See also
-
class
nengo.dists.CosineSimilarity(dimensions)[source]¶ Distribution of the cosine of the angle between two random vectors.
The “cosine similarity” is the cosine of the angle between two vectors, which is equal to the dot product of the vectors, divided by the L2-norms of the individual vectors. When these vectors are unit length, this is then simply the distribution of their dot product.
This is also equivalent to the distribution of a single coefficient from a unit vector (a single dimension of
UniformHypersphere(surface=True)). Furthermore,CosineSimilarity(d+2)is equivalent to the distribution of a single coordinate from points uniformly sampled from the d-dimensional unit ball (a single dimension ofUniformHypersphere(surface=False).sample(n, d)). These relationships have been detailed in [Voelker2017].This can be used to calculate an intercept
c = ppf(1 - p)such thatdot(u, v) >= cwith probabilityp, for random unit vectorsuandv. In other words, a neuron with interceptppf(1 - p)will fire with probabilitypfor a random unit length input.- Parameters
- dimensions: int
Dimensionality of the complete unit vector.
See also
-
sample(self, n, d=None, rng=numpy.random)[source]¶ Samples the distribution.
- Parameters
- nint
Number samples to take.
- dint or None, optional
The number of dimensions to return. If this is an int, the return value will be of shape
(n, d). If None, the return value will be of shape(n,).- rng
numpy.random.RandomState, optional Random number generator state.
- Returns
- samples(n,) or (n, d) array_like
Samples as a 1d or 2d array depending on
d. The second dimension enumerates the dimensions of the process.
-
cdf(x)[source]¶ Cumulative distribution function.
Note
Requires SciPy.
- Parameters
- xarray_like
Evaluation points in [0, 1].
- Returns
- cdfarray_like
Probability that
X <= x.
Learning rule types¶
Base class for all learning rule objects. |
|
Prescribed Error Sensitivity learning rule. |
|
Recursive least-squares rule for online decoder optimization. |
|
Bienenstock-Cooper-Munroe learning rule. |
|
Oja learning rule. |
|
Vector Oja learning rule. |
-
class
nengo.learning_rules.LearningRuleTypeSizeInParam(name, default=Unconfigurable, low=None, high=None, low_open=False, high_open=False, optional=False, readonly=None)[source]¶
-
class
nengo.learning_rules.LearningRuleType(learning_rate=Default<1e-06>, size_in=0)[source]¶ Base class for all learning rule objects.
To use a learning rule, pass it as a
learning_rule_typekeyword argument to theConnectionon which you want to do learning.Each learning rule exposes two important pieces of metadata that the builder uses to determine what information should be stored.
The
size_inis the dimensionality of the incoming error signal. It can either take an integer or one of the following string values:'pre': vector error signal in pre-object space'post': vector error signal in post-object space'mid': vector error signal in theconn.size_midspace'pre_state': vector error signal in pre-synaptic ensemble space'post_state': vector error signal in post-synaptic ensemble space
The difference between
'post_state'and'post'is that with the former, if aNeuronsobject is passed, it will use the dimensionality of the correspondingEnsemble, whereas the latter simply uses thepostobjectsize_in. Similarly with'pre_state'and'pre'.The
modifiesattribute denotes the signal targeted by the rule. Options are:'encoders''decoders''weights'
- Parameters
- learning_ratefloat, optional
A scalar indicating the rate at which
modifieswill be adjusted.- size_inint, str, optional
Dimensionality of the error signal (see above).
- Attributes
- learning_ratefloat
A scalar indicating the rate at which
modifieswill be adjusted.- size_inint, str
Dimensionality of the error signal.
- modifiesstr
The signal targeted by the learning rule.
-
class
nengo.PES(learning_rate=Default<0.0001>, pre_synapse=Default<Lowpass(tau=0.005)>)[source]¶ Prescribed Error Sensitivity learning rule.
Modifies a connection’s decoders to minimize an error signal provided through a connection to the connection’s learning rule.
- Parameters
- learning_ratefloat, optional
A scalar indicating the rate at which weights will be adjusted.
- pre_synapse
Synapse, optional Synapse model used to filter the pre-synaptic activities.
- Attributes
- learning_ratefloat
A scalar indicating the rate at which weights will be adjusted.
- pre_synapse
Synapse Synapse model used to filter the pre-synaptic activities.
-
class
nengo.RLS(learning_rate=Default<0.001>, pre_synapse=Default<Lowpass(tau=0.005)>)[source]¶ Recursive least-squares rule for online decoder optimization.
This implements an online version of the standard least-squares solvers used to learn connection weights offline (e.g.
nengo.solvers.LstsqL2). It can be applied in the same scenarios asPES, to minimize an error signal.The cost of RLS is \(\mathcal{O}(n^2)\) extra time and memory. If possible, it is more efficient to do the learning offline using e.g.
LstsqL2.- Parameters
- learning_ratefloat, optional
Effective learning rate. This is better understood as \(\frac{1}{\alpha}\), where \(\alpha\) is an L2-regularization term. A large learning rate means little regularization, which implies quick over-fitting. A small learning rate means large regularization, which translates to slower learning 2.
- pre_synapseSynapse, optional
Synapse model applied to the pre-synaptic neural activities.
See also
Notes
RLS works by maintaining the inverse neural correlation matrix, \(P = \Gamma^{-1}\), where \(\Gamma = A^T A + \alpha I\) are the regularized correlations, \(A\) is a matrix of (possibly filtered) neural activities, and \(\alpha\) is an L2-regularization term controlled by the
learning_rate. \(P\) is used to project the error signal and update the weights each time-step.References
- 2
Sussillo, D., & Abbott, L. F. (2009). Generating coherent patterns of activity from chaotic neural networks. Neuron, 63(4), 544-557.
Examples
Below, we compare
PESagainstRLS, learning a feed-forward communication channel (identity function) online, starting with 100 spiking LIF neurons with decoders (weights) set to zero. A faster learning rate forPESresults in over-fitting to the most recent online example, while a slower learning rate does not learn quickly enough. This is a general problem with greedy optimization.RLSperforms better since it is L2-optimal.from nengo.learning_rules import PES, RLS tau = 0.005 learning_rules = ( PES(learning_rate=1e-3, pre_synapse=tau), RLS(learning_rate=1e-3, pre_synapse=tau), ) with nengo.Network() as model: u = nengo.Node(output=lambda t: np.sin(2 * np.pi * t)) probes = [] for lr in learning_rules: e = nengo.Node(size_in=1, output=lambda t, e: e if t < 1 else 0) x = nengo.Ensemble(100, 1, seed=0) y = nengo.Node(size_in=1) nengo.Connection(u, e, synapse=None, transform=-1) nengo.Connection(u, x, synapse=None) conn = nengo.Connection( x, y, synapse=None, learning_rule_type=lr, function=lambda x: 0 ) nengo.Connection(y, e, synapse=None) nengo.Connection(e, conn.learning_rule, synapse=tau) probes.append(nengo.Probe(y, synapse=tau)) probes.append(nengo.Probe(u, synapse=tau)) with nengo.Simulator(model) as sim: sim.run(2.0) plt.plot(sim.trange(), sim.data[probes[0]], label=str(learning_rules[0])) plt.plot(sim.trange(), sim.data[probes[1]], label=str(learning_rules[1])) plt.plot(sim.trange(), sim.data[probes[2]], label="Ideal", linestyle="--") plt.vlines([1], -1, 1, label="Training -> Testing") plt.ylim(-2, 2) plt.legend(loc="upper right") plt.xlabel("Time (s)")
-
class
nengo.BCM(learning_rate=Default<1e-09>, pre_synapse=Default<Lowpass(tau=0.005)>, post_synapse=Default<None>, theta_synapse=Default<Lowpass(tau=1.0)>)[source]¶ Bienenstock-Cooper-Munroe learning rule.
Modifies connection weights as a function of the presynaptic activity and the difference between the postsynaptic activity and the average postsynaptic activity.
- Parameters
- learning_ratefloat, optional
A scalar indicating the rate at which weights will be adjusted.
- pre_synapse
Synapse, optional Synapse model used to filter the pre-synaptic activities.
- post_synapse
Synapse, optional Synapse model used to filter the post-synaptic activities. If None,
post_synapsewill be the same aspre_synapse.- theta_synapse
Synapse, optional Synapse model used to filter the theta signal.
Notes
The BCM rule is dependent on pre and post neural activities, not decoded values, and so is not affected by changes in the size of pre and post ensembles. However, if you are decoding from the post ensemble, the BCM rule will have an increased effect on larger post ensembles because more connection weights are changing. In these cases, it may be advantageous to scale the learning rate on the BCM rule by
1 / post.n_neurons.- Attributes
- learning_ratefloat
A scalar indicating the rate at which weights will be adjusted.
- post_synapse
Synapse Synapse model used to filter the post-synaptic activities.
- pre_synapse
Synapse Synapse model used to filter the pre-synaptic activities.
- theta_synapse
Synapse Synapse model used to filter the theta signal.
-
class
nengo.Oja(learning_rate=Default<1e-06>, pre_synapse=Default<Lowpass(tau=0.005)>, post_synapse=Default<None>, beta=Default<1.0>)[source]¶ Oja learning rule.
Modifies connection weights according to the Hebbian Oja rule, which augments typically Hebbian coactivity with a “forgetting” term that is proportional to the weight of the connection and the square of the postsynaptic activity.
- Parameters
- learning_ratefloat, optional
A scalar indicating the rate at which weights will be adjusted.
- pre_synapse
Synapse, optional Synapse model used to filter the pre-synaptic activities.
- post_synapse
Synapse, optional Synapse model used to filter the post-synaptic activities. If None,
post_synapsewill be the same aspre_synapse.- betafloat, optional
A scalar weight on the forgetting term.
Notes
The Oja rule is dependent on pre and post neural activities, not decoded values, and so is not affected by changes in the size of pre and post ensembles. However, if you are decoding from the post ensemble, the Oja rule will have an increased effect on larger post ensembles because more connection weights are changing. In these cases, it may be advantageous to scale the learning rate on the Oja rule by
1 / post.n_neurons.
-
class
nengo.Voja(learning_rate=Default<0.01>, post_synapse=Default<Lowpass(tau=0.005)>)[source]¶ Vector Oja learning rule.
Modifies an ensemble’s encoders to be selective to its inputs.
A connection to the learning rule will provide a scalar weight for the learning rate, minus 1. For instance, 0 is normal learning, -1 is no learning, and less than -1 causes anti-learning or “forgetting”.
- Parameters
- post_taufloat, optional
Filter constant on activities of neurons in post population.
- learning_ratefloat, optional
A scalar indicating the rate at which encoders will be adjusted.
- post_synapse
Synapse, optional Synapse model used to filter the post-synaptic activities.
- Attributes
- learning_ratefloat
A scalar indicating the rate at which encoders will be adjusted.
- post_synapse
Synapse Synapse model used to filter the post-synaptic activities.
Neuron types¶
Compute firing rates (in Hz) for given vector input, |
|
Base class for Nengo neuron models. |
|
Signifies that an ensemble should simulate in direct mode. |
|
A rectified linear neuron model. |
|
A rectified integrate and fire neuron model. |
|
A non-spiking neuron model whose response curve is a sigmoid. |
|
A non-spiking neuron model whose response curve is a hyperbolic tangent. |
|
Non-spiking version of the leaky integrate-and-fire (LIF) neuron model. |
|
Spiking version of the leaky integrate-and-fire (LIF) neuron model. |
|
Adaptive non-spiking version of the LIF neuron model. |
|
Adaptive spiking version of the LIF neuron model. |
|
Izhikevich neuron model. |
|
Base class for neuron types that turn rate types into spiking ones. |
|
Turn a rate neuron type into a spiking one with regular inter-spike intervals. |
|
Turn a rate neuron type into a spiking one using stochastic rounding. |
|
Turn a rate neuron type into a spiking one with Poisson spiking statistics. |
-
nengo.neurons.settled_firingrate(step, J, state, dt=0.001, settle_time=0.1, sim_time=1.0)[source]¶ Compute firing rates (in Hz) for given vector input,
x.Unlike the default naive implementation, this approach takes into account some characteristics of spiking neurons. We start by simulating the neurons for a short amount of time, to let any initial transients settle. Then, we run the neurons for a second and find the average (which should approximate the firing rate).
- Parameters
- stepfunction
the step function of the neuron type
- Jndarray
a vector of currents to generate firing rates from
- statedict of ndarrays
additional state needed by the step function
-
class
nengo.neurons.NeuronType(initial_state=None)[source]¶ Base class for Nengo neuron models.
- Parameters
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
- Attributes
- state{str: Distribution}
State variables held by the neuron type during simulation. Values in the dict indicate their initial values, or how to obtain those initial values. These elements can also be probed in the neuron population.
- negativebool
Whether the neurons can emit negative outputs (i.e. negative spikes or rates).
-
current(x, gain, bias)[source]¶ Compute current injected in each neuron given input, gain and bias.
Note that
xis assumed to be already projected onto the encoders associated with the neurons and normalized to radius 1, so the maximum expected current for a neuron occurs when input for that neuron is 1.- Parameters
- x(n_samples,) or (n_samples, n_neurons) array_like
Scalar inputs for which to calculate current.
- gain(n_neurons,) array_like
Gains associated with each neuron.
- bias(n_neurons,) array_like
Bias current associated with each neuron.
- Returns
- current(n_samples, n_neurons)
Current to be injected in each neuron.
-
gain_bias(max_rates, intercepts)[source]¶ Compute the gain and bias needed to satisfy max_rates, intercepts.
This takes the neurons, approximates their response function, and then uses that approximation to find the gain and bias value that will give the requested intercepts and max_rates.
Note that this default implementation is very slow! Whenever possible, subclasses should override this with a neuron-specific implementation.
- Parameters
- max_rates(n_neurons,) array_like
Maximum firing rates of neurons.
- intercepts(n_neurons,) array_like
X-intercepts of neurons.
- Returns
- gain(n_neurons,) array_like
Gain associated with each neuron. Sometimes denoted alpha.
- bias(n_neurons,) array_like
Bias current associated with each neuron.
-
max_rates_intercepts(gain, bias)[source]¶ Compute the max_rates and intercepts given gain and bias.
Note that this default implementation is very slow! Whenever possible, subclasses should override this with a neuron-specific implementation.
- Parameters
- gain(n_neurons,) array_like
Gain associated with each neuron. Sometimes denoted alpha.
- bias(n_neurons,) array_like
Bias current associated with each neuron.
- Returns
- max_rates(n_neurons,) array_like
Maximum firing rates of neurons.
- intercepts(n_neurons,) array_like
X-intercepts of neurons.
-
rates(x, gain, bias)[source]¶ Compute firing rates (in Hz) for given input
x.This default implementation takes the naive approach of running the step function for a second. This should suffice for most rate-based neuron types; for spiking neurons it will likely fail (those models should override this function).
Note that
xis assumed to be already projected onto the encoders associated with the neurons and normalized to radius 1, so the maximum expected rate for a neuron occurs when input for that neuron is 1.- Parameters
- x(n_samples,) or (n_samples, n_neurons) array_like
Scalar inputs for which to calculate rates.
- gain(n_neurons,) array_like
Gains associated with each neuron.
- bias(n_neurons,) array_like
Bias current associated with each neuron.
- Returns
- rates(n_samples, n_neurons) ndarray
The firing rates at each given value of
x.
-
step(dt, J, output, **state)[source]¶ Implements the differential equation for this neuron type.
At a minimum, NeuronType subclasses must implement this method. That implementation should modify the
outputparameter rather than returning anything, for efficiency reasons.- Parameters
- dtfloat
Simulation timestep.
- J(n_neurons,) array_like
Input currents associated with each neuron.
- output(n_neurons,) array_like
Output activity associated with each neuron (e.g., spikes or firing rates).
- state{str: array_like}
State variables associated with the population.
-
class
nengo.neurons.NeuronTypeParam(name, default=Unconfigurable, optional=False, readonly=None)[source]¶
-
class
nengo.Direct(initial_state=None)[source]¶ Signifies that an ensemble should simulate in direct mode.
In direct mode, the ensemble represents and transforms signals perfectly, rather than through a neural approximation. Note that direct mode ensembles with recurrent connections can easily diverge; most other neuron types will instead saturate at a certain high firing rate.
-
class
nengo.RectifiedLinear(amplitude=1, initial_state=None)[source]¶ A rectified linear neuron model.
Each neuron is modeled as a rectified line. That is, the neuron’s activity scales linearly with current, unless it passes below zero, at which point the neural activity will stay at zero.
- Parameters
- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
-
class
nengo.SpikingRectifiedLinear(amplitude=1, initial_state=None)[source]¶ A rectified integrate and fire neuron model.
Each neuron is modeled as a rectified line. That is, the neuron’s activity scales linearly with current, unless the current is less than zero, at which point the neural activity will stay at zero. This is a spiking version of the RectifiedLinear neuron model.
- Parameters
- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output spikes of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
-
class
nengo.Sigmoid(tau_ref=0.0025, initial_state=None)[source]¶ A non-spiking neuron model whose response curve is a sigmoid.
Since the tuning curves are strictly positive, the
interceptscorrespond to the inflection point of each sigmoid. That is,f(intercept) = 0.5wherefis the pure sigmoid function.- Parameters
- tau_reffloat
The neuron refractory period, in seconds. The maximum firing rate of the neurons is
1 / tau_ref. Must be positive (i.e.tau_ref > 0).- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
-
class
nengo.Tanh(tau_ref=0.0025, initial_state=None)[source]¶ A non-spiking neuron model whose response curve is a hyperbolic tangent.
- Parameters
- tau_reffloat
The neuron refractory period, in seconds. The maximum firing rate of the neurons is
1 / tau_ref. Must be positive (i.e.tau_ref > 0).- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
-
class
nengo.LIFRate(tau_rc=0.02, tau_ref=0.002, amplitude=1, initial_state=None)[source]¶ Non-spiking version of the leaky integrate-and-fire (LIF) neuron model.
- Parameters
- tau_rcfloat
Membrane RC time constant, in seconds. Affects how quickly the membrane voltage decays to zero in the absence of input (larger = slower decay).
- tau_reffloat
Absolute refractory period, in seconds. This is how long the membrane voltage is held at zero after a spike.
- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output spikes of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
-
class
nengo.LIF(tau_rc=0.02, tau_ref=0.002, min_voltage=0, amplitude=1, initial_state=None)[source]¶ Spiking version of the leaky integrate-and-fire (LIF) neuron model.
- Parameters
- tau_rcfloat
Membrane RC time constant, in seconds. Affects how quickly the membrane voltage decays to zero in the absence of input (larger = slower decay).
- tau_reffloat
Absolute refractory period, in seconds. This is how long the membrane voltage is held at zero after a spike.
- min_voltagefloat
Minimum value for the membrane voltage. If
-np.inf, the voltage is never clipped.- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output spikes of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
-
class
nengo.AdaptiveLIFRate(tau_n=1, inc_n=0.01, tau_rc=0.02, tau_ref=0.002, amplitude=1, initial_state=None)[source]¶ Adaptive non-spiking version of the LIF neuron model.
Works as the LIF model, except with adapation state
n, which is subtracted from the input current. Its dynamics are:tau_n dn/dt = -n
where
nis incremented byinc_nwhen the neuron spikes.- Parameters
- tau_nfloat
Adaptation time constant. Affects how quickly the adaptation state decays to zero in the absence of spikes (larger = slower decay).
- inc_nfloat
Adaptation increment. How much the adaptation state is increased after each spike.
- tau_rcfloat
Membrane RC time constant, in seconds. Affects how quickly the membrane voltage decays to zero in the absence of input (larger = slower decay).
- tau_reffloat
Absolute refractory period, in seconds. This is how long the membrane voltage is held at zero after a spike.
- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output spikes of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
References
- 1
Camera, Giancarlo La, et al. “Minimal models of adapted neuronal response to in Vivo-Like input currents.” Neural computation 16.10 (2004): 2101-2124.
-
class
nengo.AdaptiveLIF(tau_n=1, inc_n=0.01, tau_rc=0.02, tau_ref=0.002, min_voltage=0, amplitude=1, initial_state=None)[source]¶ Adaptive spiking version of the LIF neuron model.
Works as the LIF model, except with adapation state
n, which is subtracted from the input current. Its dynamics are:tau_n dn/dt = -n
where
nis incremented byinc_nwhen the neuron spikes.- Parameters
- tau_nfloat
Adaptation time constant. Affects how quickly the adaptation state decays to zero in the absence of spikes (larger = slower decay).
- inc_nfloat
Adaptation increment. How much the adaptation state is increased after each spike.
- tau_rcfloat
Membrane RC time constant, in seconds. Affects how quickly the membrane voltage decays to zero in the absence of input (larger = slower decay).
- tau_reffloat
Absolute refractory period, in seconds. This is how long the membrane voltage is held at zero after a spike.
- min_voltagefloat
Minimum value for the membrane voltage. If
-np.inf, the voltage is never clipped.- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output spikes of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
References
- 1
Camera, Giancarlo La, et al. “Minimal models of adapted neuronal response to in Vivo-Like input currents.” Neural computation 16.10 (2004): 2101-2124.
-
class
nengo.Izhikevich(tau_recovery=0.02, coupling=0.2, reset_voltage=- 65.0, reset_recovery=8.0, initial_state=None)[source]¶ Izhikevich neuron model.
This implementation is based on the original paper [1]; however, we rename some variables for clarity. What was originally ‘v’ we term ‘voltage’, which represents the membrane potential of each neuron. What was originally ‘u’ we term ‘recovery’, which represents membrane recovery, “which accounts for the activation of K+ ionic currents and inactivation of Na+ ionic currents.” The ‘a’, ‘b’, ‘c’, and ‘d’ parameters are also renamed (see the parameters below).
We use default values that correspond to regular spiking (‘RS’) neurons. For other classes of neurons, set the parameters as follows.
Intrinsically bursting (IB):
reset_voltage=-55, reset_recovery=4Chattering (CH):
reset_voltage=-50, reset_recovery=2Fast spiking (FS):
tau_recovery=0.1Low-threshold spiking (LTS):
coupling=0.25Resonator (RZ):
tau_recovery=0.1, coupling=0.26
- Parameters
- tau_recoveryfloat, optional
(Originally ‘a’) Time scale of the recovery variable.
- couplingfloat, optional
(Originally ‘b’) How sensitive recovery is to subthreshold fluctuations of voltage.
- reset_voltagefloat, optional
(Originally ‘c’) The voltage to reset to after a spike, in millivolts.
- reset_recoveryfloat, optional
(Originally ‘d’) The recovery value to reset to after a spike.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
References
- 1
E. M. Izhikevich, “Simple model of spiking neurons.” IEEE Transactions on Neural Networks, vol. 14, no. 6, pp. 1569-1572. (http://www.izhikevich.org/publications/spikes.pdf)
-
class
nengo.neurons.RatesToSpikesNeuronType(base_type, amplitude=1.0, initial_state=None)[source]¶ Base class for neuron types that turn rate types into spiking ones.
-
gain_bias(max_rates, intercepts)[source]¶ Compute the gain and bias needed to satisfy max_rates, intercepts.
This takes the neurons, approximates their response function, and then uses that approximation to find the gain and bias value that will give the requested intercepts and max_rates.
Note that this default implementation is very slow! Whenever possible, subclasses should override this with a neuron-specific implementation.
- Parameters
- max_rates(n_neurons,) array_like
Maximum firing rates of neurons.
- intercepts(n_neurons,) array_like
X-intercepts of neurons.
- Returns
- gain(n_neurons,) array_like
Gain associated with each neuron. Sometimes denoted alpha.
- bias(n_neurons,) array_like
Bias current associated with each neuron.
-
max_rates_intercepts(gain, bias)[source]¶ Compute the max_rates and intercepts given gain and bias.
Note that this default implementation is very slow! Whenever possible, subclasses should override this with a neuron-specific implementation.
- Parameters
- gain(n_neurons,) array_like
Gain associated with each neuron. Sometimes denoted alpha.
- bias(n_neurons,) array_like
Bias current associated with each neuron.
- Returns
- max_rates(n_neurons,) array_like
Maximum firing rates of neurons.
- intercepts(n_neurons,) array_like
X-intercepts of neurons.
-
rates(x, gain, bias)[source]¶ Compute firing rates (in Hz) for given input
x.This default implementation takes the naive approach of running the step function for a second. This should suffice for most rate-based neuron types; for spiking neurons it will likely fail (those models should override this function).
Note that
xis assumed to be already projected onto the encoders associated with the neurons and normalized to radius 1, so the maximum expected rate for a neuron occurs when input for that neuron is 1.- Parameters
- x(n_samples,) or (n_samples, n_neurons) array_like
Scalar inputs for which to calculate rates.
- gain(n_neurons,) array_like
Gains associated with each neuron.
- bias(n_neurons,) array_like
Bias current associated with each neuron.
- Returns
- rates(n_samples, n_neurons) ndarray
The firing rates at each given value of
x.
-
step(dt, J, output, **state)[source]¶ Implements the differential equation for this neuron type.
At a minimum, NeuronType subclasses must implement this method. That implementation should modify the
outputparameter rather than returning anything, for efficiency reasons.- Parameters
- dtfloat
Simulation timestep.
- J(n_neurons,) array_like
Input currents associated with each neuron.
- output(n_neurons,) array_like
Output activity associated with each neuron (e.g., spikes or firing rates).
- state{str: array_like}
State variables associated with the population.
-
-
class
nengo.RegularSpiking(base_type, amplitude=1.0, initial_state=None)[source]¶ Turn a rate neuron type into a spiking one with regular inter-spike intervals.
Spikes at regular intervals based on the rates of the base neuron type. [1]
- Parameters
- base_typeNeuronType
A rate-based neuron type to convert to a regularly spiking neuron.
- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output spikes of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
References
- 1
Voelker, A. R., Rasmussen, D., & Eliasmith, C. (2020). A Spike in Performance: Training Hybrid-Spiking Neural Networks with Quantized Activation Functions. arXiv preprint arXiv:2002.03553. (https://export.arxiv.org/abs/2002.03553)
-
step(dt, J, output, voltage)[source]¶ Implements the differential equation for this neuron type.
At a minimum, NeuronType subclasses must implement this method. That implementation should modify the
outputparameter rather than returning anything, for efficiency reasons.- Parameters
- dtfloat
Simulation timestep.
- J(n_neurons,) array_like
Input currents associated with each neuron.
- output(n_neurons,) array_like
Output activity associated with each neuron (e.g., spikes or firing rates).
- state{str: array_like}
State variables associated with the population.
-
class
nengo.StochasticSpiking(base_type, amplitude=1.0, initial_state=None)[source]¶ Turn a rate neuron type into a spiking one using stochastic rounding.
The expected number of spikes per timestep
e = dt * ris determined by the base type firing raterand the timestepdt. Given the fractional partfand integer partqofe, the number of generated spikes isqwith probability1 - fandq + 1with probabilityf. Foremuch less than one, this is very similar to Poisson statistics.- Parameters
- base_typeNeuronType
A rate-based neuron type to convert to a stochastic spiking neuron.
- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output spikes of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
-
step(dt, J, output, rng, **base_state)[source]¶ Implements the differential equation for this neuron type.
At a minimum, NeuronType subclasses must implement this method. That implementation should modify the
outputparameter rather than returning anything, for efficiency reasons.- Parameters
- dtfloat
Simulation timestep.
- J(n_neurons,) array_like
Input currents associated with each neuron.
- output(n_neurons,) array_like
Output activity associated with each neuron (e.g., spikes or firing rates).
- state{str: array_like}
State variables associated with the population.
-
class
nengo.PoissonSpiking(base_type, amplitude=1.0, initial_state=None)[source]¶ Turn a rate neuron type into a spiking one with Poisson spiking statistics.
Spikes with Poisson probability based on the rates of the base neuron type.
- Parameters
- base_typeNeuronType
A rate-based neuron type to convert to a Poisson spiking neuron.
- amplitudefloat
Scaling factor on the neuron output. Corresponds to the relative amplitude of the output spikes of the neuron.
- initial_state{str: Distribution or array_like}
Mapping from state variables names to their desired initial value. These values will override the defaults set in the class’s state attribute.
-
step(dt, J, output, rng, **base_state)[source]¶ Implements the differential equation for this neuron type.
At a minimum, NeuronType subclasses must implement this method. That implementation should modify the
outputparameter rather than returning anything, for efficiency reasons.- Parameters
- dtfloat
Simulation timestep.
- J(n_neurons,) array_like
Input currents associated with each neuron.
- output(n_neurons,) array_like
Output activity associated with each neuron (e.g., spikes or firing rates).
- state{str: array_like}
State variables associated with the population.
Processes¶
A general system with input, output, and state. |
|
Full-spectrum white noise process. |
|
Filtered white noise process. |
|
Brown noise process (aka Brownian noise, red noise, Wiener process). |
|
An ideal low-pass filtered white noise process. |
|
Present a series of inputs, each for the same fixed length of time. |
|
A piecewise function with different options for interpolation. |
-
class
nengo.processes.WhiteNoise(dist=Gaussian(mean=0, std=1), scale=True, **kwargs)[source]¶ Full-spectrum white noise process.
- Parameters
- distDistribution, optional
The distribution from which to draw samples.
- scalebool, optional
Whether to scale the white noise for integration. Integrating white noise requires using a time constant of
sqrt(dt)instead ofdton the noise term [1], to ensure the magnitude of the integrated noise does not change withdt.- seedint, optional
Random number seed. Ensures noise will be the same each run.
References
- 1
Gillespie, D.T. (1996) Exact numerical simulation of the Ornstein- Uhlenbeck process and its integral. Phys. Rev. E 54, pp. 2084-91.
-
make_step(shape_in, shape_out, dt, rng, state)[source]¶ Create function that advances the process forward one time step.
This must be implemented by all custom processes. The parameters below indicate what information is provided by the builder.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- rng
numpy.random.RandomState A random number generator.
- state{string:
numpy.ndarray} A dictionary mapping keys to signals, where the signals fully represent the state of the process. The signals are initialized by
Process.make_state.New in version 3.0.0.
-
class
nengo.processes.FilteredNoise(synapse=Lowpass(tau=0.005), dist=Gaussian(mean=0, std=1), scale=True, **kwargs)[source]¶ Filtered white noise process.
This process takes white noise and filters it using the provided synapse.
- Parameters
- synapseSynapse, optional
The synapse to use to filter the noise.
- distDistribution, optional
The distribution used to generate the white noise.
- scalebool, optional
Whether to scale the white noise for integration, making the output signal invariant to
dt.- seedint, optional
Random number seed. Ensures noise will be the same each run.
-
make_state(shape_in, shape_out, dt, dtype=None)[source]¶ Get a dictionary of signals to represent the state of this process.
The builder uses this to allocate memory for the process state, so that the state can be represented as part of the whole simulator state.
New in version 3.0.0.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- dtype
numpy.dtype The data type requested by the builder. If
None, then this function is free to choose the best type for the signals involved.
- Returns
- initial_state{string:
numpy.ndarray} A dictionary mapping keys to arrays containing the initial state values. The keys will be used to identify the signals in
Process.make_step.
- initial_state{string:
-
make_step(shape_in, shape_out, dt, rng, state)[source]¶ Create function that advances the process forward one time step.
This must be implemented by all custom processes. The parameters below indicate what information is provided by the builder.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- rng
numpy.random.RandomState A random number generator.
- state{string:
numpy.ndarray} A dictionary mapping keys to signals, where the signals fully represent the state of the process. The signals are initialized by
Process.make_state.New in version 3.0.0.
-
class
nengo.processes.BrownNoise(dist=Gaussian(mean=0, std=1), **kwargs)[source]¶ Brown noise process (aka Brownian noise, red noise, Wiener process).
This process is the integral of white noise.
- Parameters
- distDistribution, optional
The distribution used to generate the white noise.
- seedint, optional
Random number seed. Ensures noise will be the same each run.
-
class
nengo.processes.WhiteSignal(period, high, rms=0.5, y0=None, **kwargs)[source]¶ An ideal low-pass filtered white noise process.
This signal is created in the frequency domain, and designed to have exactly equal power at all frequencies below the cut-off frequency, and no power above the cut-off.
The signal is naturally periodic, so it can be used beyond its period while still being continuous with continuous derivatives.
- Parameters
- periodfloat
A white noise signal with this period will be generated. Samples will repeat after this duration.
- highfloat
The cut-off frequency of the low-pass filter, in Hz. Must not exceed the Nyquist frequency for the simulation timestep, which is
0.5 / dt.- rmsfloat, optional
The root mean square power of the filtered signal
- y0float, optional
Align the phase of each output dimension to begin at the value that is closest (in absolute value) to y0.
- seedint, optional
Random number seed. Ensures noise will be the same each run.
-
make_step(shape_in, shape_out, dt, rng, state)[source]¶ Create function that advances the process forward one time step.
This must be implemented by all custom processes. The parameters below indicate what information is provided by the builder.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- rng
numpy.random.RandomState A random number generator.
- state{string:
numpy.ndarray} A dictionary mapping keys to signals, where the signals fully represent the state of the process. The signals are initialized by
Process.make_state.New in version 3.0.0.
-
class
nengo.processes.PresentInput(inputs, presentation_time, **kwargs)[source]¶ Present a series of inputs, each for the same fixed length of time.
- Parameters
- inputsarray_like
Inputs to present, where each row is an input. Rows will be flattened.
- presentation_timefloat
Show each input for this amount of time (in seconds).
-
make_step(shape_in, shape_out, dt, rng, state)[source]¶ Create function that advances the process forward one time step.
This must be implemented by all custom processes. The parameters below indicate what information is provided by the builder.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- rng
numpy.random.RandomState A random number generator.
- state{string:
numpy.ndarray} A dictionary mapping keys to signals, where the signals fully represent the state of the process. The signals are initialized by
Process.make_state.New in version 3.0.0.
-
class
nengo.processes.PiecewiseDataParam(name, default=Unconfigurable, optional=False, readonly=None)[source]¶ Piecewise-specific validation for the data dictionary.
In the
Piecewisedata dict, the keys are points in time (float) and values are numerical constants or callables of the same dimensionality.
-
class
nengo.processes.Piecewise(data, interpolation='zero', **kwargs)[source]¶ A piecewise function with different options for interpolation.
Given an input dictionary of
{0: 0, 0.5: -1, 0.75: 0.5, 1: 0}, this process will emit the numerical values (0, -1, 0.5, 0) starting at the corresponding time points (0, 0.5, 0.75, 1).The keys in the input dictionary must be times (float or int). The values in the dictionary can be floats, lists of floats, or numpy arrays. All lists or numpy arrays must be of the same length, as the output shape of the process will be determined by the shape of the values.
Interpolation on the data points using
scipy.interpolateis also supported. The default interpolation is ‘zero’, which creates a piecewise function whose values change at the specified time points. So the above example would be shortcut for:def function(t): if t < 0.5: return 0 elif t < 0.75: return -1 elif t < 1: return 0.5 else: return 0
For times before the first specified time, an array of zeros (of the correct length) will be emitted. This means that the above can be simplified to:
from nengo.processes import Piecewise Piecewise({0.5: -1, 0.75: 0.5, 1: 0})
- Parameters
- datadict
A dictionary mapping times to the values that should be emitted at those times. Times must be numbers (ints or floats), while values can be numbers, lists of numbers, numpy arrays of numbers, or callables that return any of those options.
- interpolationstr, optional
One of ‘linear’, ‘nearest’, ‘slinear’, ‘quadratic’, ‘cubic’, or ‘zero’. Specifies how to interpolate between times with specified value. ‘zero’ creates a plain piecewise function whose values begin at corresponding time points, while all other options interpolate as described in
scipy.interpolate.
Examples
from nengo.processes import Piecewise process = Piecewise({0.5: 1, 0.75: -1, 1: 0}) with nengo.Network() as model: u = nengo.Node(process, size_out=process.default_size_out) up = nengo.Probe(u) with nengo.Simulator(model, progress_bar=False) as sim: sim.run(1.5) f = sim.data[up] t = sim.trange() print(f[t == 0.2]) print(f[t == 0.58])
[[ 0.]] [[ 1.]]
- Attributes
- datadict
A dictionary mapping times to the values that should be emitted at those times. Times are numbers (ints or floats), while values can be numbers, lists of numbers, numpy arrays of numbers, or callables that return any of those options.
- interpolationstr
One of ‘linear’, ‘nearest’, ‘slinear’, ‘quadratic’, ‘cubic’, or ‘zero’. Specifies how to interpolate between times with specified value. ‘zero’ creates a plain piecewise function whose values change at corresponding time points, while all other options interpolate as described in
scipy.interpolate.
-
make_step(shape_in, shape_out, dt, rng, state)[source]¶ Create function that advances the process forward one time step.
This must be implemented by all custom processes. The parameters below indicate what information is provided by the builder.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- rng
numpy.random.RandomState A random number generator.
- state{string:
numpy.ndarray} A dictionary mapping keys to signals, where the signals fully represent the state of the process. The signals are initialized by
Process.make_state.New in version 3.0.0.
-
class
nengo.Process(default_size_in=0, default_size_out=1, default_dt=0.001, seed=None)[source]¶ A general system with input, output, and state.
For more details on how to use processes and make custom process subclasses, see Processes and how to use them.
- Parameters
- default_size_inint
Sets the default size in for nodes using this process.
- default_size_outint
Sets the default size out for nodes running this process. Also, if
dis not specified inrunorrun_steps, this will be used.- default_dtfloat
If
dtis not specified inrun,run_steps,ntrange, ortrange, this will be used.- seedint, optional
Random number seed. Ensures random factors will be the same each run.
- Attributes
- default_dtfloat
If
dtis not specified inrun,run_steps,ntrange, ortrange, this will be used.- default_size_inint
The default size in for nodes using this process.
- default_size_outint
The default size out for nodes running this process. Also, if
dis not specified inrunorrun_steps, this will be used.- seedint or None
Random number seed. Ensures random factors will be the same each run.
-
apply(self, x, d=None, dt=None, rng=numpy.random, copy=True, **kwargs)[source]¶ Run process on a given input.
Keyword arguments that do not appear in the parameter list below will be passed to the
make_stepfunction of this process.- Parameters
- xndarray
The input signal given to the process.
- dint, optional
Output dimensionality. If None,
default_size_outwill be used.- dtfloat, optional
Simulation timestep. If None,
default_dtwill be used.- rng
numpy.random.RandomState Random number generator used for stochstic processes.
- copybool, optional
If True, a new output array will be created for output. If False, the input signal
xwill be overwritten.
-
get_rng(rng)[source]¶ Get a properly seeded independent RNG for the process step.
- Parameters
- rng
numpy.random.RandomState The parent random number generator to use if the seed is not set.
- rng
-
make_state(shape_in, shape_out, dt, dtype=None)[source]¶ Get a dictionary of signals to represent the state of this process.
The builder uses this to allocate memory for the process state, so that the state can be represented as part of the whole simulator state.
New in version 3.0.0.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- dtype
numpy.dtype The data type requested by the builder. If
None, then this function is free to choose the best type for the signals involved.
- Returns
- initial_state{string:
numpy.ndarray} A dictionary mapping keys to arrays containing the initial state values. The keys will be used to identify the signals in
Process.make_step.
- initial_state{string:
-
make_step(shape_in, shape_out, dt, rng, state)[source]¶ Create function that advances the process forward one time step.
This must be implemented by all custom processes. The parameters below indicate what information is provided by the builder.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- rng
numpy.random.RandomState A random number generator.
- state{string:
numpy.ndarray} A dictionary mapping keys to signals, where the signals fully represent the state of the process. The signals are initialized by
Process.make_state.New in version 3.0.0.
-
run(self, t, d=None, dt=None, rng=numpy.random, **kwargs)[source]¶ Run process without input for given length of time.
Keyword arguments that do not appear in the parameter list below will be passed to the
make_stepfunction of this process.- Parameters
- tfloat
The length of time to run.
- dint, optional
Output dimensionality. If None,
default_size_outwill be used.- dtfloat, optional
Simulation timestep. If None,
default_dtwill be used.- rng
numpy.random.RandomState Random number generator used for stochstic processes.
-
run_steps(self, n_steps, d=None, dt=None, rng=numpy.random, **kwargs)[source]¶ Run process without input for given number of steps.
Keyword arguments that do not appear in the parameter list below will be passed to the
make_stepfunction of this process.- Parameters
- n_stepsint
The number of steps to run.
- dint, optional
Output dimensionality. If None,
default_size_outwill be used.- dtfloat, optional
Simulation timestep. If None,
default_dtwill be used.- rng
numpy.random.RandomState Random number generator used for stochstic processes.
Solvers¶
Classes concerned with solving for decoders or full weight matrices.
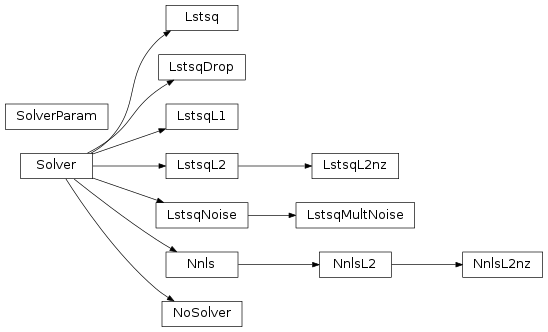
Decoder or weight solver. |
|
Unregularized least-squares solver. |
|
Least-squares solver with additive Gaussian white noise. |
|
Least-squares solver with multiplicative white noise. |
|
Least-squares solver with L2 regularization. |
|
Least-squares solver with L2 regularization on non-zero components. |
|
Least-squares solver with L1 and L2 regularization (elastic net). |
|
Find sparser decoders/weights by dropping small values. |
|
Non-negative least-squares solver without regularization. |
|
Non-negative least-squares solver with L2 regularization. |
|
Non-negative least-squares with L2 regularization on nonzero components. |
|
Manually pass in weights, bypassing the decoder solver. |
-
class
nengo.solvers.Solver(weights=False)[source]¶ Decoder or weight solver.
A solver can have the
weightsparameter equal toTrueorFalse.Weight solvers are used to form neuron-to-neuron weight matrices. They can be compositional or non-compositional. Non-compositional solvers must operate on the whole neuron-to-neuron weight matrix (i.e., each target is a separate postsynaptic current, without the bias term), while compositional solvers operate in the decoded state-space (i.e., each target is a dimension in state-space). Compositional solvers then combine the returned
Xwith the transform and/or encoders to generate the full weight matrix.For a solver to be compositional, the following property must be true:
X = solver(A, Y) if and only if L(X) = solver(A, L(Y))
where
Lis some arbitrary linear operator (i.e., the transform and/or encoders for the postsynaptic population). This property can then be leveraged by the backend for efficiency. See the solver’scompositionalclass attribute to determine if it is compositional.Non-weight solvers always operate in the decoded state-space regardless of whether they are compositional or non-compositional.
-
class
nengo.solvers.SolverParam(name, default=Unconfigurable, optional=False, readonly=None)[source]¶ A parameter in which the value is a
Solverinstance.
-
class
nengo.solvers.Lstsq(weights=False, rcond=0.01)[source]¶ Unregularized least-squares solver.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- rcondfloat, optional
Cut-off ratio for small singular values (see
numpy.linalg.lstsq).
- Attributes
- rcondfloat
Cut-off ratio for small singular values (see
numpy.linalg.lstsq).- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.LstsqNoise(weights=False, noise=0.1, solver=Cholesky())[source]¶ Least-squares solver with additive Gaussian white noise.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- noisefloat, optional
Amount of noise, as a fraction of the neuron activity.
- solver
LeastSquaresSolver, optional Subsolver to use for solving the least squares problem.
- Attributes
- noisefloat
Amount of noise, as a fraction of the neuron activity.
- solver
LeastSquaresSolver Subsolver to use for solving the least squares problem.
- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.LstsqMultNoise(weights=False, noise=0.1, solver=Cholesky())[source]¶ Least-squares solver with multiplicative white noise.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- noisefloat, optional
Amount of noise, as a fraction of the neuron activity.
- solver
LeastSquaresSolver, optional Subsolver to use for solving the least squares problem.
- Attributes
- noisefloat
Amount of noise, as a fraction of the neuron activity.
- solver
LeastSquaresSolver Subsolver to use for solving the least squares problem.
- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.LstsqL2(weights=False, reg=0.1, solver=Cholesky())[source]¶ Least-squares solver with L2 regularization.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- regfloat, optional
Amount of regularization, as a fraction of the neuron activity.
- solver
LeastSquaresSolver, optional Subsolver to use for solving the least squares problem.
- Attributes
- regfloat
Amount of regularization, as a fraction of the neuron activity.
- solver
LeastSquaresSolver Subsolver to use for solving the least squares problem.
- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.LstsqL2nz(weights=False, reg=0.1, solver=Cholesky())[source]¶ Least-squares solver with L2 regularization on non-zero components.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- regfloat, optional
Amount of regularization, as a fraction of the neuron activity.
- solver
LeastSquaresSolver, optional Subsolver to use for solving the least squares problem.
- Attributes
- regfloat
Amount of regularization, as a fraction of the neuron activity.
- solver
LeastSquaresSolver Subsolver to use for solving the least squares problem.
- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.LstsqL1(weights=False, l1=0.0001, l2=1e-06, max_iter=1000)[source]¶ Least-squares solver with L1 and L2 regularization (elastic net).
This method is well suited for creating sparse decoders or weight matrices.
Note
Requires scikit-learn.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- l1float, optional
Amount of L1 regularization.
- l2float, optional
Amount of L2 regularization.
- max_iterint, optional
Maximum number of iterations for the underlying elastic net.
- Attributes
- l1float
Amount of L1 regularization.
- l2float
Amount of L2 regularization.
- weightsbool
If False, solve for decoders. If True, solve for weights.
- max_iterint
Maximum number of iterations for the underlying elastic net.
-
class
nengo.solvers.LstsqDrop(weights=False, drop=0.25, solver1=LstsqL2(reg=0.001), solver2=LstsqL2())[source]¶ Find sparser decoders/weights by dropping small values.
This solver first solves for coefficients (decoders/weights) with L2 regularization, drops those nearest to zero, and retrains remaining.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- dropfloat, optional
Fraction of decoders or weights to set to zero.
- solver1Solver, optional
Solver for finding the initial decoders.
- solver2Solver, optional
Used for re-solving for the decoders after dropout.
- Attributes
- dropfloat
Fraction of decoders or weights to set to zero.
- solver1Solver
Solver for finding the initial decoders.
- solver2Solver
Used for re-solving for the decoders after dropout.
- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.Nnls(weights=False)[source]¶ Non-negative least-squares solver without regularization.
Similar to
Lstsq, except the output values are non-negative.If solving for non-negative weights, it is important that the intercepts of the post-population are also non-negative, since neurons with negative intercepts will never be silent, affecting output accuracy.
Note
Requires SciPy.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- Attributes
- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.NnlsL2(weights=False, reg=0.1)[source]¶ Non-negative least-squares solver with L2 regularization.
Similar to
LstsqL2, except the output values are non-negative.If solving for non-negative weights, it is important that the intercepts of the post-population are also non-negative, since neurons with negative intercepts will never be silent, affecting output accuracy.
Note
Requires SciPy.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- regfloat, optional
Amount of regularization, as a fraction of the neuron activity.
- Attributes
- regfloat
Amount of regularization, as a fraction of the neuron activity.
- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.NnlsL2nz(weights=False, reg=0.1)[source]¶ Non-negative least-squares with L2 regularization on nonzero components.
Similar to
LstsqL2nz, except the output values are non-negative.If solving for non-negative weights, it is important that the intercepts of the post-population are also non-negative, since neurons with negative intercepts will never be silent, affecting output accuracy.
Note
Requires SciPy.
- Parameters
- weightsbool, optional
If False, solve for decoders. If True, solve for weights.
- regfloat, optional
Amount of regularization, as a fraction of the neuron activity.
- Attributes
- regfloat
Amount of regularization, as a fraction of the neuron activity.
- weightsbool
If False, solve for decoders. If True, solve for weights.
-
class
nengo.solvers.NoSolver(values=None, weights=False)[source]¶ Manually pass in weights, bypassing the decoder solver.
- Parameters
- values(n_neurons, size_out) array_like, optional
The array of decoders to use.
size_outis the dimensionality of the decoded signal (determined by the connection function). IfNone, which is the default, the solver will return an appropriately sized array of zeros.- weightsbool, optional
If False, connection will use factored weights (decoders from this solver, transform, and encoders). If True, connection will use a full weight matrix (created by linearly combining decoder, transform, and encoders).
- Attributes
- values(n_neurons, size_out) array_like, optional
The array of decoders to use.
size_outis the dimensionality of the decoded signal (determined by the connection function). IfNone, which is the default, the solver will return an appropriately sized array of zeros.- weightsbool, optional
If False, connection will use factored weights (decoders from this solver, transform, and encoders). If True, connection will use a full weight matrix (created by linearly combining decoder, transform, and encoders).
Solver methods¶
These solvers are to be passed as arguments to Solver objects.
For example:
from nengo.solvers import LstsqL2
from nengo.utils.least_squares_solvers import SVD
with nengo.Network():
ens_a = nengo.Ensemble(10, 1)
ens_b = nengo.Ensemble(10, 1)
nengo.Connection(ens_a, ens_b, solver=LstsqL2(solver=SVD()))
Extract data from A/Y matrices. |
|
Returns the root-mean-squared error (RMSE) of the solution X. |
|
Linear least squares system solver. |
|
Solve a least-squares system using the Cholesky decomposition. |
|
Solve a least-squares system using Scipy’s conjugate gradient. |
|
Solve a least-squares system using Scipy’s LSMR. |
|
Solve a least-squares system using conjugate gradient. |
|
Solve a multiple-RHS least-squares system using block conj. |
|
Solve a least-squares system using full SVD. |
|
Solve a least-squares system using a randomized (partial) SVD. |
|
A parameter where the value is a LeastSquaresSolver. |
-
nengo.utils.least_squares_solvers.rmses(A, X, Y)[source]¶ Returns the root-mean-squared error (RMSE) of the solution X.
-
class
nengo.utils.least_squares_solvers.LeastSquaresSolver[source]¶ Linear least squares system solver.
-
class
nengo.utils.least_squares_solvers.Cholesky(transpose=None)[source]¶ Solve a least-squares system using the Cholesky decomposition.
-
class
nengo.utils.least_squares_solvers.ConjgradScipy(tol=0.0001, atol=1e-08)[source]¶ Solve a least-squares system using Scipy’s conjugate gradient.
- Parameters
References
- 1(1,2)
scipy.sparse.linalg.cg documentation, https://docs.scipy.org/doc/scipy/reference/generated/scipy.sparse.linalg.cg.html
-
class
nengo.utils.least_squares_solvers.LSMRScipy(tol=0.0001)[source]¶ Solve a least-squares system using Scipy’s LSMR.
-
class
nengo.utils.least_squares_solvers.Conjgrad(tol=0.01, maxiters=None, X0=None)[source]¶ Solve a least-squares system using conjugate gradient.
-
class
nengo.utils.least_squares_solvers.BlockConjgrad(tol=0.01, X0=None)[source]¶ Solve a multiple-RHS least-squares system using block conj. gradient.
-
class
nengo.utils.least_squares_solvers.RandomizedSVD(n_components=60, n_oversamples=10, n_iter=0)[source]¶ Solve a least-squares system using a randomized (partial) SVD.
Useful for solving large matrices quickly, but non-optimally.
- Parameters
- n_componentsint, optional
The number of SVD components to compute. A small survey of activity matrices suggests that the first 60 components capture almost all the variance.
- n_oversamplesint, optional
The number of additional samples on the range of A.
- n_iterint, optional
The number of power iterations to perform (can help with noisy data).
See also
sklearn.utils.extmath.randomized_svdFunction used by this class
Synapse models¶
Abstract base class for synapse models. |
|
General linear time-invariant (LTI) system synapse. |
|
Standard first-order lowpass filter synapse. |
|
Alpha-function filter synapse. |
|
Triangular finite impulse response (FIR) synapse. |
-
class
nengo.synapses.Synapse(default_size_in=1, default_size_out=None, default_dt=0.001, seed=None)[source]¶ Abstract base class for synapse models.
Conceptually, a synapse model emulates a biological synapse, taking in input in the form of released neurotransmitter and opening ion channels to allow more or less current to flow into the neuron.
In Nengo, the implementation of a synapse is as a specific case of a
Processin which the input and output shapes are the same. The input is the current across the synapse, and the output is the current that will be induced in the postsynaptic neuron.Synapses also contain the
Synapse.filtandSynapse.filtfiltmethods, which make it easy to use Nengo’s synapse models outside of Nengo simulations.- Parameters
- default_size_inint, optional
The size_in used if not specified.
- default_size_outint
The size_out used if not specified. If None, will be the same as default_size_in.
- default_dtfloat
The simulation timestep used if not specified.
- seedint, optional
Random number seed. Ensures random factors will be the same each run.
- Attributes
- default_dtfloat
The simulation timestep used if not specified.
- default_size_inint
The size_in used if not specified.
- default_size_outint
The size_out used if not specified.
- seedint, optional
Random number seed. Ensures random factors will be the same each run.
-
make_state(shape_in, shape_out, dt, dtype=None, y0=None)[source]¶ Get a dictionary of signals to represent the state of this process.
The builder uses this to allocate memory for the process state, so that the state can be represented as part of the whole simulator state.
New in version 3.0.0.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- dtype
numpy.dtype The data type requested by the builder. If
None, then this function is free to choose the best type for the signals involved.
- Returns
- initial_state{string:
numpy.ndarray} A dictionary mapping keys to arrays containing the initial state values. The keys will be used to identify the signals in
Process.make_step.
- initial_state{string:
-
filt(x, dt=None, axis=0, y0=0, copy=True, filtfilt=False)[source]¶ Filter
xwith this synapse model.- Parameters
- xarray_like
The signal to filter.
- dtfloat, optional
The timestep of the input signal. If None,
default_dtwill be used.- axisint, optional
The axis along which to filter.
- y0array_like, optional
The starting state of the filter output. Must be zero for unstable linear systems.
- copybool, optional
Whether to copy the input data, or simply work in-place.
- filtfiltbool, optional
If True, runs the process forward then backward on the signal, for zero-phase filtering (like Matlab’s
filtfilt).
-
filtfilt(x, **kwargs)[source]¶ Zero-phase filtering of
xusing this filter.Equivalent to
filt(x, filtfilt=True, **kwargs).
-
class
nengo.LinearFilter(num, den, analog=True, method='zoh', **kwargs)[source]¶ General linear time-invariant (LTI) system synapse.
This class can be used to implement any linear filter, given the filter’s transfer function. [1]
- Parameters
- numarray_like
Numerator coefficients of transfer function.
- denarray_like
Denominator coefficients of transfer function.
- analogboolean, optional
Whether the synapse coefficients are analog (i.e. continuous-time), or discrete. Analog coefficients will be converted to discrete for simulation using the simulator
dt.- methodstring
The method to use for discretization (if
analogis True). Seescipy.signal.cont2discretefor information about the options.New in version 3.0.0.
References
- Attributes
- analogboolean
Whether the synapse coefficients are analog (i.e. continuous-time), or discrete. Analog coefficients will be converted to discrete for simulation using the simulator
dt.- denndarray
Denominator coefficients of transfer function.
- numndarray
Numerator coefficients of transfer function.
- methodstring
The method to use for discretization (if
analogis True). Seescipy.signal.cont2discretefor information about the options.
-
evaluate(frequencies)[source]¶ Evaluate the transfer function at the given frequencies.
Examples
Using the
evaluatefunction to make a Bode plot:import matplotlib.pyplot as plt synapse = nengo.synapses.LinearFilter([1], [0.02, 1]) f = np.logspace(-1, 3, 100) y = synapse.evaluate(f) plt.subplot(211); plt.semilogx(f, 20*np.log10(np.abs(y))) plt.xlabel('frequency [Hz]'); plt.ylabel('magnitude [dB]') plt.subplot(212); plt.semilogx(f, np.angle(y)) plt.xlabel('frequency [Hz]'); plt.ylabel('phase [radians]')
-
make_state(shape_in, shape_out, dt, dtype=None, y0=0)[source]¶ Get a dictionary of signals to represent the state of this process.
The builder uses this to allocate memory for the process state, so that the state can be represented as part of the whole simulator state.
New in version 3.0.0.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- dtype
numpy.dtype The data type requested by the builder. If
None, then this function is free to choose the best type for the signals involved.
- Returns
- initial_state{string:
numpy.ndarray} A dictionary mapping keys to arrays containing the initial state values. The keys will be used to identify the signals in
Process.make_step.
- initial_state{string:
-
make_step(shape_in, shape_out, dt, rng, state)[source]¶ Returns a
Stepinstance that implements the linear filter.
-
class
OneXScalar(A, B, C, D, X)[source]¶ Step for systems with one state element, no passthrough, and a size-1 input.
Using the builtin float math improves performance.
-
class
nengo.Lowpass(tau, **kwargs)[source]¶ Standard first-order lowpass filter synapse.
The impulse-response function is given by:
f(t) = (1 / tau) * exp(-t / tau)
- Parameters
- taufloat
The time constant of the filter in seconds.
- Attributes
- taufloat
The time constant of the filter in seconds.
-
class
nengo.Alpha(tau, **kwargs)[source]¶ Alpha-function filter synapse.
The impulse-response function is given by:
alpha(t) = (t / tau**2) * exp(-t / tau)
and was found by [1] to be a good basic model for synapses.
- Parameters
- taufloat
The time constant of the filter in seconds.
References
- 1
Mainen, Z.F. and Sejnowski, T.J. (1995). Reliability of spike timing in neocortical neurons. Science (New York, NY), 268(5216):1503-6.
- Attributes
- taufloat
The time constant of the filter in seconds.
-
class
nengo.synapses.Triangle(t, **kwargs)[source]¶ Triangular finite impulse response (FIR) synapse.
This synapse has a triangular and finite impulse response. The length of the triangle is
tseconds; thus the digital filter will havet / dt + 1taps.- Parameters
- tfloat
Length of the triangle, in seconds.
- Attributes
- tfloat
Length of the triangle, in seconds.
-
make_state(shape_in, shape_out, dt, dtype=None, y0=0)[source]¶ Get a dictionary of signals to represent the state of this process.
The builder uses this to allocate memory for the process state, so that the state can be represented as part of the whole simulator state.
New in version 3.0.0.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- dtype
numpy.dtype The data type requested by the builder. If
None, then this function is free to choose the best type for the signals involved.
- Returns
- initial_state{string:
numpy.ndarray} A dictionary mapping keys to arrays containing the initial state values. The keys will be used to identify the signals in
Process.make_step.
- initial_state{string:
-
make_step(shape_in, shape_out, dt, rng, state)[source]¶ Create function that advances the process forward one time step.
This must be implemented by all custom processes. The parameters below indicate what information is provided by the builder.
- Parameters
- shape_intuple
The shape of the input signal.
- shape_outtuple
The shape of the output signal.
- dtfloat
The simulation timestep.
- rng
numpy.random.RandomState A random number generator.
- state{string:
numpy.ndarray} A dictionary mapping keys to signals, where the signals fully represent the state of the process. The signals are initialized by
Process.make_state.New in version 3.0.0.
Transforms¶
A base class for connection transforms. |
|
A dense matrix transformation between an input and output signal. |
|
Represents a sparse matrix. |
|
A sparse matrix transformation between an input and output signal. |
|
An N-dimensional convolutional transform. |
|
An N-dimensional transposed convolutional transform. |
|
Represents shape information with variable channel position. |
|
Directly pass the signal through without any transform operations. |
|
alias of |
|
alias of |
-
class
nengo.transforms.Transform[source]¶ A base class for connection transforms.
New in version 3.0.0.
-
sample(self, rng=numpy.random)[source]¶ Returns concrete weights to implement the specified transform.
- Parameters
- rng
numpy.random.RandomState, optional Random number generator state.
- rng
- Returns
- array_like
Transform weights
-
property
size_in¶ Expected size of input to transform.
-
property
size_out¶ Expected size of output from transform.
-
-
class
nengo.transforms.ChannelShapeParam(name, default=Unconfigurable, length=None, low=0, optional=False, readonly=None)[source]¶ A parameter where the value must be a shape with channels.
New in version 3.0.0.
-
class
nengo.Dense(shape, init=1.0)[source]¶ A dense matrix transformation between an input and output signal.
New in version 3.0.0.
- Parameters
- shapetuple of int
The shape of the dense matrix:
(size_out, size_in).- init
Distributionor array_like, optional A Distribution used to initialize the transform matrix, or a concrete instantiation for the matrix. If the matrix is square we also allow a scalar (equivalent to
np.eye(n) * init) or a vector (equivalent tonp.diag(init)) to represent the matrix more compactly.
-
sample(self, rng=numpy.random)[source]¶ Returns concrete weights to implement the specified transform.
- Parameters
- rng
numpy.random.RandomState, optional Random number generator state.
- rng
- Returns
- array_like
Transform weights
-
property
init_shape¶ The shape of the initial value.
-
property
size_in¶ Expected size of input to transform.
-
property
size_out¶ Expected size of output from transform.
-
class
nengo.transforms.SparseInitParam(name, default=Unconfigurable, optional=False, readonly=None)[source]¶
-
class
nengo.transforms.SparseMatrix(indices, data, shape)[source]¶ Represents a sparse matrix.
New in version 3.0.0.
- Parameters
- indicesarray_like of int
An Nx2 array of integers indicating the (row,col) coordinates for the N non-zero elements in the matrix.
- dataarray_like or
Distribution An Nx1 array defining the value of the nonzero elements in the matrix (corresponding to
indices), or aDistributionthat will be used to initialize the nonzero elements.- shapetuple of int
Shape of the full matrix.
-
allocate()[source]¶ Return a
scipy.sparse.csr_matrixor dense matrix equivalent.We mark this data as readonly to be consistent with how other data associated with signals are allocated. If this allocated data is to be modified, it should be copied first.
-
sample(self, rng=numpy.random)[source]¶ Convert
Distributiondata to fixed array.- Parameters
- rng
numpy.random.RandomState Random number generator that will be used when sampling distribution.
- rng
- Returns
- matrix
SparseMatrix A new
SparseMatrixinstance withDistributionconverted to array ifself.datais aDistribution, otherwise simply returnsself.
- matrix
-
class
nengo.Sparse(shape, indices=None, init=1.0)[source]¶ A sparse matrix transformation between an input and output signal.
New in version 3.0.0.
- Parameters
- shapetuple of int
The full shape of the sparse matrix:
(size_out, size_in).- indicesarray_like of int
An Nx2 array of integers indicating the (row,col) coordinates for the N non-zero elements in the matrix.
- init
Distributionor array_like, optional A Distribution used to initialize the transform matrix, or a concrete instantiation for the matrix. If the matrix is square we also allow a scalar (equivalent to
np.eye(n) * init) or a vector (equivalent tonp.diag(init)) to represent the matrix more compactly.
-
sample(self, rng=numpy.random)[source]¶ Returns concrete weights to implement the specified transform.
- Parameters
- rng
numpy.random.RandomState, optional Random number generator state.
- rng
- Returns
- array_like
Transform weights
-
property
size_in¶ Expected size of input to transform.
-
property
size_out¶ Expected size of output from transform.
-
class
nengo.Convolution(n_filters, input_shape, kernel_size=(3, 3), strides=(1, 1), padding='valid', channels_last=True, init=Uniform(low=- 1, high=1))[source]¶ An N-dimensional convolutional transform.
The dimensionality of the convolution is determined by the input shape.
New in version 3.0.0.
- Parameters
- n_filtersint
The number of convolutional filters to apply.
- input_shapetuple of int or
ChannelShape Shape of the input signal to the convolution; e.g.,
(height, width, channels)for a 2D convolution withchannels_last=True.- kernel_sizetuple of int, optional
Size of the convolutional kernels (1 element for a 1D convolution, 2 for a 2D convolution, etc.).
- stridestuple of int, optional
Stride of the convolution (1 element for a 1D convolution, 2 for a 2D convolution, etc.).
- padding
"same"or"valid", optional Padding method for input signal. “Valid” means no padding, and convolution will only be applied to the fully-overlapping areas of the input signal (meaning the output will be smaller). “Same” means that the input signal is zero-padded so that the output is the same shape as the input.
- channels_lastbool, optional
If
True(default), the channels are the last dimension in the input signal (e.g., a 28x28 image with 3 channels would have shape(28, 28, 3)).Falsemeans that channels are the first dimension (e.g.,(3, 28, 28)).- init
Distributionorndarray, optional A predefined kernel with shape
kernel_size + (input_channels, n_filters), or aDistributionthat will be used to initialize the kernel.
Notes
As is typical in neural networks, this is technically correlation rather than convolution (because the kernel is not flipped).
-
property
output_shape¶ Output shape after applying convolution to input.
-
class
nengo.ConvolutionTranspose(n_filters, input_shape, output_shape=None, kernel_size=(3, 3), strides=(1, 1), padding='valid', channels_last=True, init=Uniform(low=- 1, high=1))[source]¶ An N-dimensional transposed convolutional transform.
This performs the transpose operation of
Convolution. Thekernel_size,strides, andpaddingparameters all act as inConvolution, so this transform will be the transpose of aConvolutiontransform with those parameters. Then_filtersandinput_shapeparameters are relative to this transform. The output shape is ambiguous, and can thus be specified (i.e. withConvolution, there can be more than one input shape that produces the same output shape, so here, there are multiple valid output shapes for some input shapes).The dimensionality of the transpose convolution is determined by the input shape.
New in version 3.2.0.
- Parameters
- n_filtersint
The number of channels in the output of this transform.
- input_shapetuple of int or
ChannelShape Shape of the input signal to this transform; e.g.,
(height, width, channels)for a 2D convolution withchannels_last=True.- output_shapetuple of int or
ChannelShape, optional Shape of the output signal of this transform; e.g.,
(output_height, output_width, n_filters)for a 2D convolution withchannels_last=True. Defaults to the smallest valid output shape.- kernel_sizetuple of int, optional
Size of the convolutional kernels (1 element for a 1D convolution, 2 for a 2D convolution, etc.).
- stridestuple of int, optional
Stride of the convolution (1 element for a 1D convolution, 2 for a 2D convolution, etc.).
- padding
"same"or"valid", optional Padding method for corresponding
Convolution.- channels_lastbool, optional
If
True(default), the channels are the last dimension in the input signal (e.g., a 28x28 image with 3 channels would have shape(28, 28, 3)).Falsemeans that channels are the first dimension (e.g.,(3, 28, 28)).- init
Distributionorndarray, optional A predefined kernel with shape
kernel_size + (input_channels, n_filters), or aDistributionthat will be used to initialize the kernel.
Notes
As is typical in neural networks, this is technically correlation rather than convolution (because the kernel is not flipped).
-
class
nengo.transforms.ChannelShape(shape, channels_last=True)[source]¶ Represents shape information with variable channel position.
New in version 3.0.0.
- Parameters
- shapeiterable of int
Signal shape
- channels_lastbool, optional
If True (default), the last item in
shaperepresents the channels, and the rest are spatial dimensions. Otherwise, the first item inshapeis the channel dimension.
-
classmethod
from_space_and_channels(spatial_shape, n_channels, channels_last=True)[source]¶ Create a ChannelShape from a spatial shape and number of channels.
New in version 3.2.0.
- Parameters
- spatial_shapeiterable of int
The spatial part of the shape (not including channels).
- n_channelsint
The number of channels.
- channels_lastbool, optional
If True (default), the last item in
shaperepresents the channels, and the rest are spatial dimensions. Otherwise, the first item inshapeis the channel dimension.
-
property
spatial_shape¶ The spatial part of the shape (omitting channels).
-
property
size¶ The total number of elements in the represented signal.
-
property
n_channels¶ The number of channels in the represented signal.
-
property
dimensions¶ The spatial dimensionality of the represented signal.
-
class
nengo.transforms.NoTransform(size_in)[source]¶ Directly pass the signal through without any transform operations.
New in version 3.1.0.
- Parameters
- size_inint
Dimensionality of transform input and output.
-
sample(self, rng=numpy.random)[source]¶ Returns concrete weights to implement the specified transform.
- Parameters
- rng
numpy.random.RandomState, optional Random number generator state.
- rng
- Raises
- TypeError
There is nothing to sample for NoTransform, so it is an error if this is called.
-
property
size_in¶ Expected size of input to transform.
-
property
size_out¶ Expected size of output from transform.
-
nengo.Conv[source]¶ alias of
nengo.Convolution
-
nengo.ConvTranspose[source]¶ alias of
nengo.ConvolutionTranspose